News
Here you will find all the newsletters. If you would like to receive newsletters automatically, create an account and accept to receive newsletters. Create an account

Qlucore Newsletter: Reducing the cost to Analyze Cancer of Unknown Primary (CUP)
Health Evidence and Innovation at Alberta Health Services recently reported in Cancer Investigation on the diagnostic costs associated with Cancer of ...
Read more
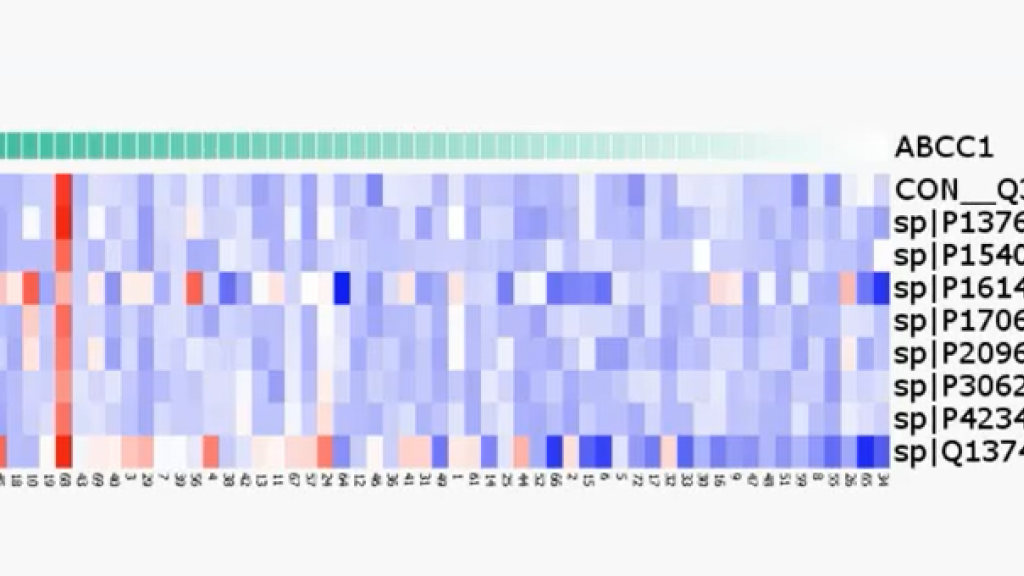
Qlucore Newsletter – Multi-Omics Analyses made Easy
With the Biomarker Workbench in Qlucore Omics Explorer, it becomes fast and straightforward to calculate correlation values and assess their statistic...
Read more

Qlucore Newsletter. New features - Easy analysis of Olink data
The latest version of Qlucore Omics Explorer includes the necessary tools and methods to efficiently import, normalize and analyze data from various O...
Read more

Qlucore Newsletter – Validated Whole Transcriptome Sequencing for diagnostics grade Gene Fusion Detection
Whole Transcriptome Sequencing is a valid method for diagnostic grade detection of gene fusions according to study in British Journal of Medicine
Read more

Qlucore Newsletter: Qlucore Advances Transcriptomic Based Cancer Diagnostics with €2.5 million in EU Support
Qlucore accelerates innovation in RNA-based clinical diagnostics, focusing on tests for acute myeloid leukemia (AML) and bladder cancer.
Read more
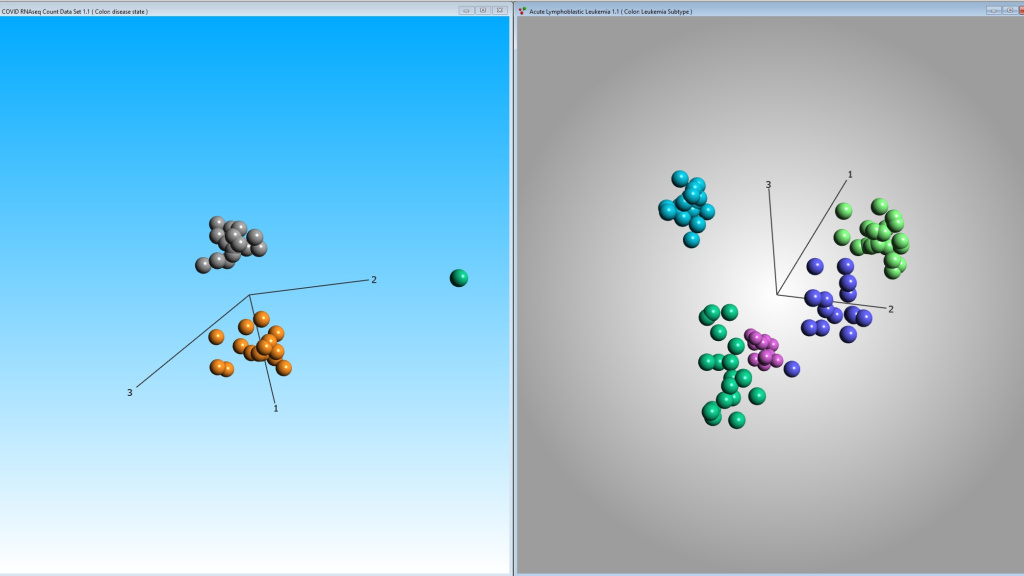
Version 3.11 is launched
Qlucore Omics Explorer (QOE) version 3.11 is launched. This release places special focus on tools and functionality that make proteomics analysis easi...
Read more

Qlucore Newsletter: RNA Sequencing is Transforming Diagnostics
RNA sequencing (RNA-seq) is rapidly proving its value in diagnostics, offering insights into gene expression, fusions, and mutations.
Read more

Qlucore Newsletter: Early Subtyping can Reduce Unclassified Lung Tumors by 80%
Unclassified tumors delay analysis and decision-making.
Read more

Qlucore Newsletter: Smarter Subtyping of Lung Cancer Samples
Looking for smarter lung cancer sample analysis?
Read more

Qlucore Newsletter: Find Patterns and Insights in Your Data
Heatmaps are powerful visualizations. They can help you uncover hidden patterns and insights in your data.
Read more

Qlucore Newsletter: Free Trainings – Available at some of Europe´s leading Research Hubs
Get better answers from your data. Enhance your understanding and use of Qlucore Omics Explorer.
Read more

Qlucore Newsletter: Let Your Data Speak with Clarity and Impact
Thousands of researchers have already benefited from the easy-to-use Qlucore Omics Explorer software, to accelerate their path to publication. Are you...