Newsletter: Easier import of RNA-seq single cell data
We are proud to introduce an upgraded version of Qlucore Omics Explorer 3.8 (3.8.15) supporting single cell data import based on the H5 format.
The key feature of the upgrade is the new single cell data import template which supports the widely used H5 format. It loads data generated with the 10X Genomics Cell Ranger pipeline. The new template also includes functionality for automatic data filtering and optional data normalization.
You find the Qlucore Template Browser with all available templates in the Launch part of the GUI. Additionally, a number of minor issues have been resolved.
Compared to version 3.7, the Qlucore Omics Explorer 3.8 adds the following functionality:
- Improved single cell data import with H5 data format support and automatic filtering
- A powerful and flexible Biomarker workbench optimized for experiments and studies in the areas of drug development and biomarker discovery.
- Expanded statistics module with easy set-up of Two-way ANOVA.
- Pie chart plot – making it possible to visualize the distribution of samples.
- ChIP seq and ATAC seq full workflow support.
- Flexible and fast Genome browser, i.e.. It is now possible to label, color and order sample annotations in two levels to add variables to a list with the mouse tool.
- Fusion gene visualization with in-depth information about breakpoints and full sequence
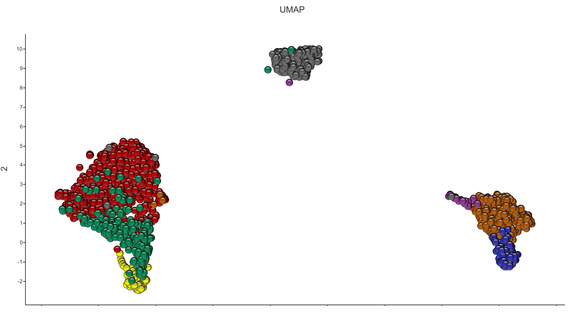
UPCOMING FREE WEBINAR:
“RNA-seq and single cell data analysis"
November 16th, 2022
15:00 GMT (+1)
Learn more about Single cell analysis in video.